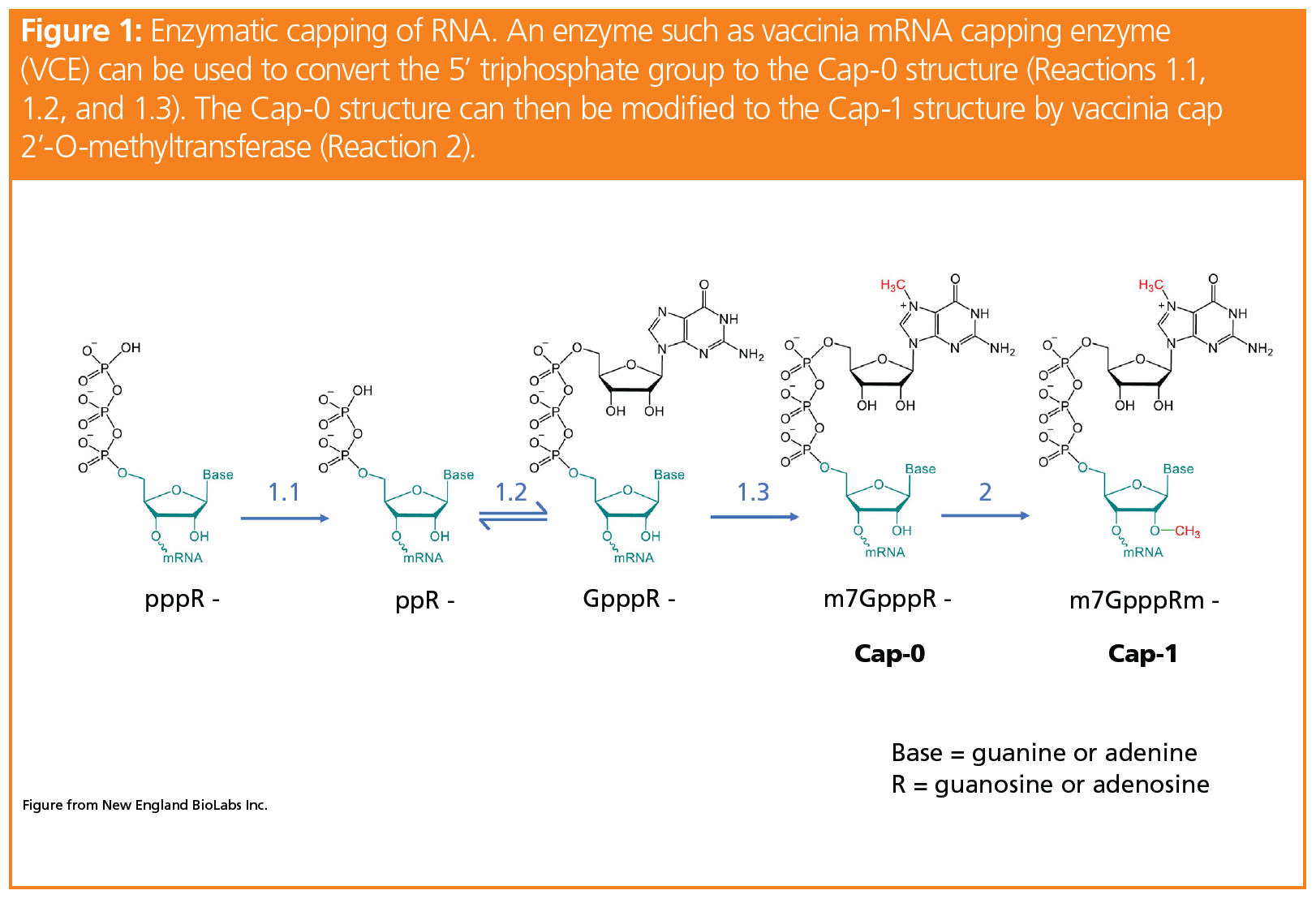
Enzymatic Assays to Explore Viral mRNA Capping Machinery - Kasprzyk - 2021 - ChemBioChem - Wiley Online Library
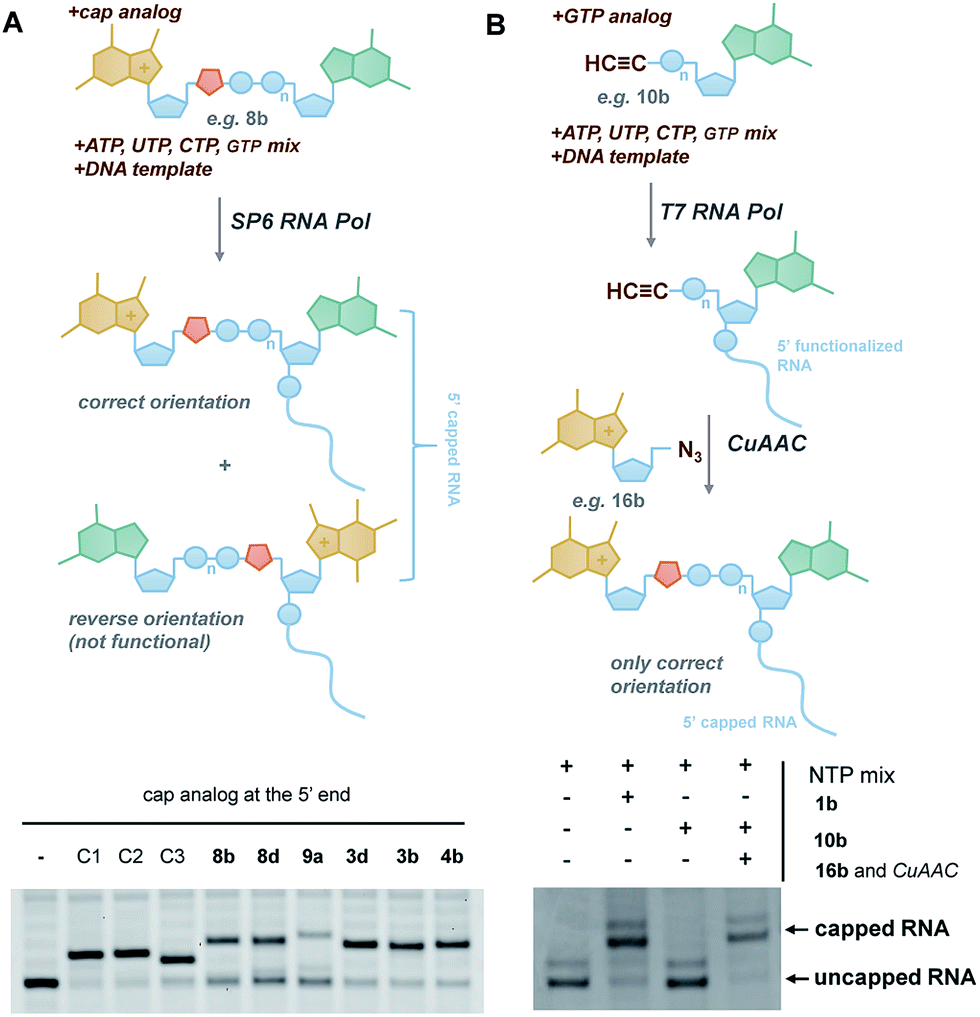
A novel route for preparing 5′ cap mimics and capped RNAs: phosphate-modified cap analogues obtained via click chemistry - Chemical Science (RSC Publishing) DOI:10.1039/C6SC02437H

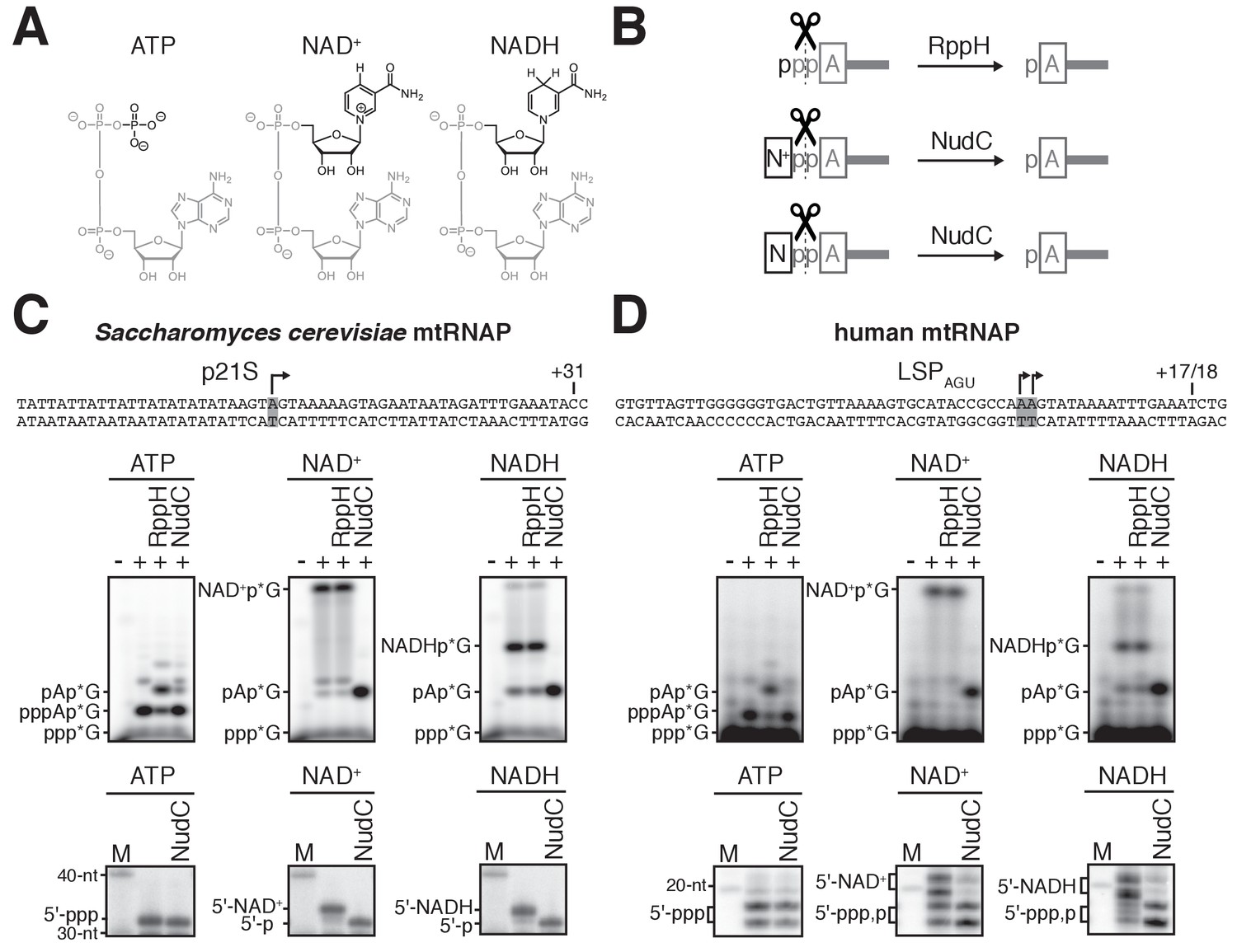
CapZyme-Seq Comprehensively Defines Promoter-Sequence Determinants for RNA 5′ Capping with NAD+ - ScienceDirect

Large-scale preparation of short, capped RNAs. (A) A longer RNA (30 nt)... | Download Scientific Diagram

Differential Inhibition of mRNA Degradation Pathways by Novel Cap Analogs* - Journal of Biological Chemistry

Pharmaceutics | Free Full-Text | Ribozyme Assays to Quantify the Capping Efficiency of In Vitro-Transcribed mRNA

Enzymatic Assays to Explore Viral mRNA Capping Machinery - Kasprzyk - 2021 - ChemBioChem - Wiley Online Library

A Mammalian Pre-mRNA 5′ End Capping Quality Control Mechanism and an Unexpected Link of Capping to Pre-mRNA Processing - ScienceDirect

Enzymatic Assays to Explore Viral mRNA Capping Machinery - Kasprzyk - 2021 - ChemBioChem - Wiley Online Library
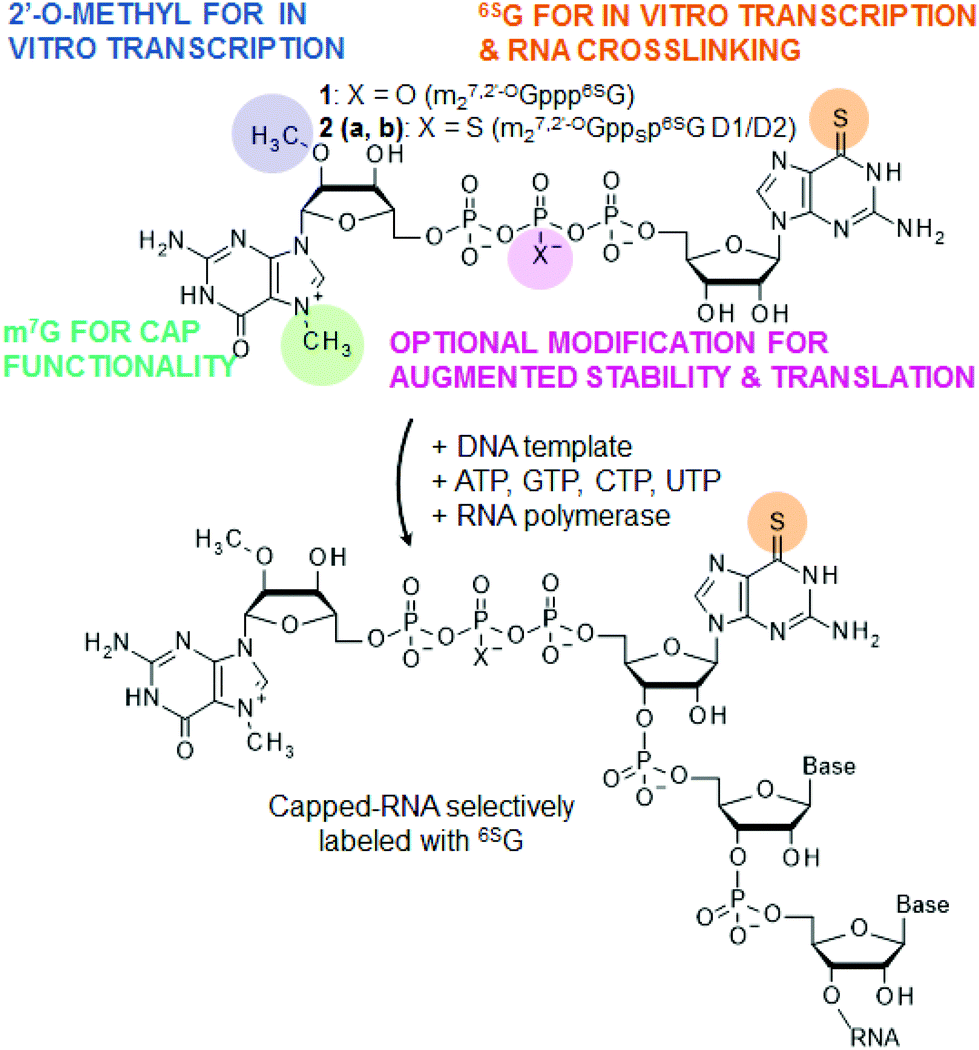
Cap analogs containing 6-thioguanosine – reagents for the synthesis of mRNAs selectively photo-crosslinkable with cap-binding biomolecules - Organic & Biomolecular Chemistry (RSC Publishing) DOI:10.1039/C4OB00059E

In vitro Transcription and Capping of Gaussia Luciferase mRNA Followed by HeLa Cell Transfection | Protocol

CAP-MAP: cap analysis protocol with minimal analyte processing, a rapid and sensitive approach to analysing mRNA cap structures | Open Biology

CAP-MAP: cap analysis protocol with minimal analyte processing, a rapid and sensitive approach to analysing mRNA cap structures | Open Biology

Pharmaceutics | Free Full-Text | Ribozyme Assays to Quantify the Capping Efficiency of In Vitro-Transcribed mRNA